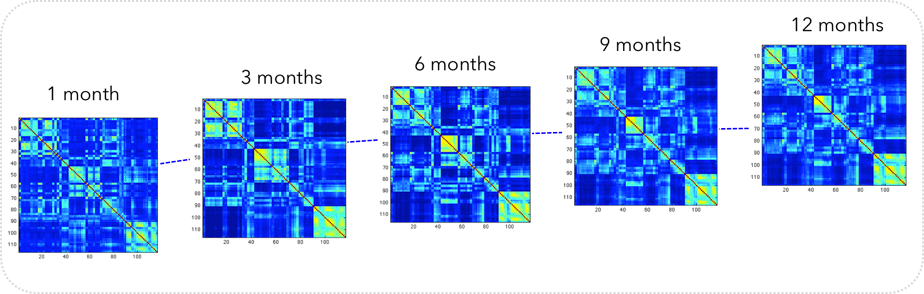
Infant brain development
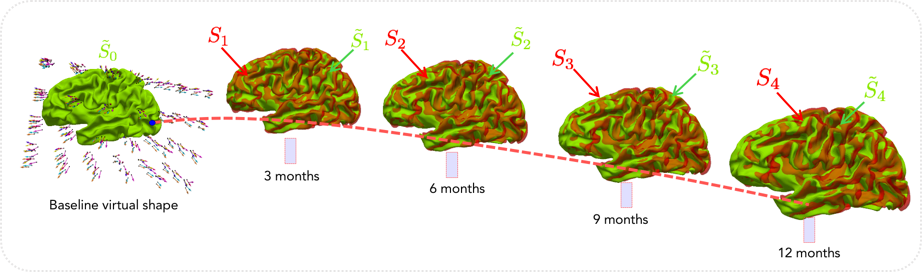
Infant brain development prediction from baseline
Motivation: Learning to predict 4D brain development from baseline brain shape (i.e. neonatal timepoint) can help predict biomarkers for neurodevelopmental disorders.
Innovation: Devising learning-based models for predicting (e.g., cortical surface, cortical surface and white matter fiber tracts, MR image) development from baseline.
Brain connectomics
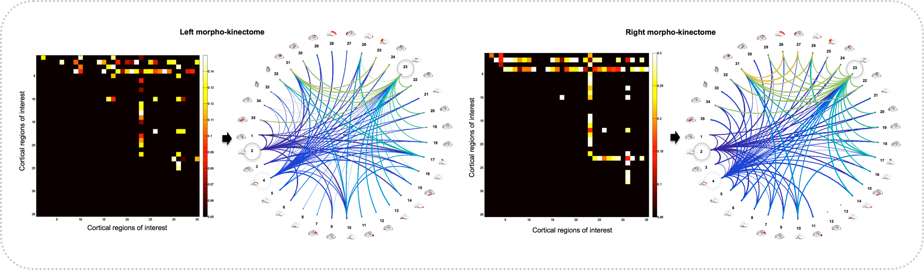
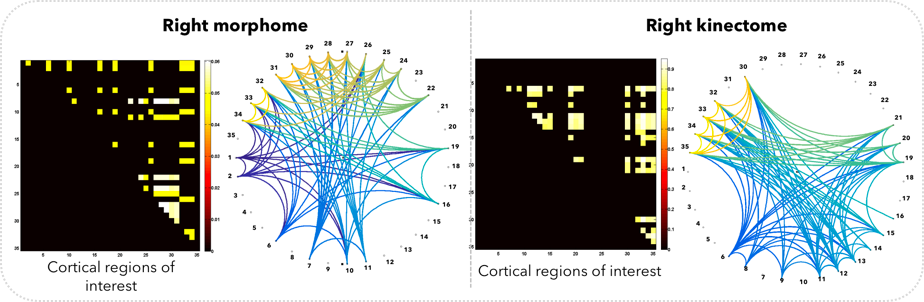
Shape-to-shape brain connectivity network (morphome) and growth-to-growth connectivity network (kinectome)
Shape-growth brain connectivity networks: morpho-kinectomes
Motivation: The development of in vivo brain connectomics field has heavily relied on using functional and diffusion Magnetic Resonance Imaging (MRI) modalities, which have fundamentally focused on studying the functional and structural relationships between pairs of anatomical regions in the brain. However, works on brain morphological (i.e., shape-to-shape) connections, which can be derived from T1 and T2 MR images, in both typical and atypical development or ageing are almost absent. Furthermore, the brain cannot be only regarded as a static shape, since it is a dynamic complex system that changes at functional, structural and morphological levels. Hence, examining the ‘connection’ between brain shape and its changes with time (e.g., growth or atrophy) may help advance our understanding of brain dynamics as well as disorders that may affect it.
Innovation: We introduce in this work three population-based shape-growth connectivity analysis tools that further extend the field of connectomics to brain morphology and dynamics: the morphome, kinectome and morpho-kinectome.
Brain network atlas estimation